Abstract
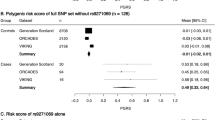
The aetiology of multiple sclerosis (MS) is uncertain. There is strong circumstantial evidence to indicate it is an autoimmune complex trait1–3. Risks for first degree relatives are increased some 20 fold over the general population4. Twin studies have shown monozygotic concordance rates of 25–30% compared to 4% for dizygotic twins and siblings5,6. Studies of adoptees and half sibs show that familial risk is determined by genes, but environmental factors strongly influence observed geographic differences3,7–9. Studies of candidate genes have been largely unrewarding10–12. We report a genome search using 257 microsatellite markers with average spacing of 15.2 cM in 100 sibling pairs (Table 1, data set 1 – DS1). A locus of λ>3 was excluded from 88% of the genome. Five loci with maximum lod scores (MLS) of > 1 were identified on chromosomes 2, 3, 5, 11 and X. Two additional data sets containing 44 (Table 1, DS2) and 78 sib pairs (Table 1, DS3) respectively, were used to further evaluate the HLA region on 6p21 and a locus on chromosome 5 with an MLS of 4.24. Markers within 6p21 gave MLS of 0.65 (nonsignificant, NS). However, D6S461, just outside the HLA region, showed significant evidence for linkage disequilibrium by the transmission disequilibrium test (TDT), in all three data sets (for DS1 λ2=10.8, adjusted P<0.01)(DS2 and DS3 λ2 = 10.9, P < 0.0005), suggesting a modest susceptibility locus in this region. On chromosome 5p results from all three data sets (222 sib pairs) yielded a multipoint MLS of 1.6. The results support genetic epidemiological evidence that several genes interact epistatically to determine heritable susceptibility.
This is a preview of subscription content, access via your institution
Access options
Subscribe to this journal
Receive 12 print issues and online access
$209.00 per year
only $17.42 per issue
Buy this article
- Purchase on Springer Link
- Instant access to full article PDF
Prices may be subject to local taxes which are calculated during checkout
Similar content being viewed by others
References
Ebers, G.C. & Sadovnick, A.D., Role of Genetic Factors in Multiple Sclerosis Susceptibility. J. Neuroimmunol. 54, 1–17 (1994).
Sadovnick, A.D. & Ebers, G.C. Epidemiology of MS: A critical overview. Can. J. Neurol. Sci. 20, 17–29 (1993).
Ebers, G.C., Sadovnick, A.D. & Risch, N.J. A genetic basis for familial aggregation in multiple sclerosis. Nature 377, 150–151 (1995).
Sadovnick, A.D., Baird, P.A. & Ward, R.H., Updated risks for relatives. Am. J. Med. Genet. 29, 533–541 (1988).
Ebers, G.C. et al. A population- based twin study in multiple sclerosis. New Engl. J. Med. 315, 1638–1642 (1986).
Mumford, C.J. et al. The British Isles survey of multiple sclerosis in twins. Neurology 44, 11–15 (1994).
Sadovnick, A.D., Ebers, G.C., Dyment, D., Risch, N. & the Canadian Collaborative Study Group. Evidence for the genetic basis of multiple sclerosis. Lancet (in the press).
Hammond, S.R. et al. The epidemiology of multiple sclerosis in three Australian cities, Perth, Newcastle and Hobart. Brain 111, 1–25 (1988).
Sadovnick, A.D., Ebers, G.C., Risch, N.J. & the Canadian Collaborative Study Group. Sibling Risks for Multiple Sclerosis. Neurology 46, A335 (1996).
Ebers, G.C., Paty, D.W., Stiller, C., Nelson, R. & Seland, T. HLA Typing in MS Sibling Pairs.Lancet 88-90 (1982).
Kellar-Wood, H.F. et al. Multiple sclerosis and the HLA-D region: linkage and association studies. J. Neuroimmunol. 58, 183–190 (1995).
Risch, N. Assessing the role of HLA-linked and unlinked determinants of disease. Am. J. Hum. Genet. 40, 1–14 (1987).
Risch, N. A note on multiple testing procedures in linkage analysis. Am. J. Hum. Genet. 48, 1058–1064 (1991).
Lander, E. & Kruglyak, L. Genetic dissection of complex traits: Guidelines for interpreting and reporting linkage results. Nature Genet. 11, 241–247 (1995).
Hillert, J. & Olerup, O., Sclerosis is associated with genes within or close to the HLA-DR-DQ subregion on a normal DR15, DQ6, Dw2 haplotype. Neurology 43, 163–168 (1993).
Tienari, R., Wikstrom, J., Sajantila, A., Palo, J. & Peltonen, L. Genetic susceptibility to multiple sclerosis linked to the myelin basic protein gene. Lancet 340, 987–991 (1993).
Wood, N.W., Holmans, P., Clayton, D., Robertson, N. & Compston, D.A.S. No linkage or association between multiple sclerosis and the myelin basic protein gene in affected sibling pairs. J. Neurol.Neurosurg. Psychiat. 57, 1191–1194 (1994).
Seboun, E., Robinson, M.A. & Doolittle, T.H. A susceptibility locus for multiple sclerosis is linked to the T-cell reception beta chain complex. Cell 57, 1095–1100 (1989).
Beall, S.S., Biddison, W.E., McFarlin, D.E., McFarland, H. & Hood, L.E. Susceptibility for multiple sclerosis is determined, in part, by inheritance of a 175-kb region of the TCR V beta locus and the HLA class II genes. J. Neuroimmunol. 45, 53–60 (1993).
Hashimoto, L., Mak, T. & Ebers, G.C. T-cell alpha chain polymorphism in MS. J. Neuroimmunol. 40, 41–48 (1992).
Hillert, J., Chunmoal, L. & Olerup, O. No association with germline T cell receptor σ-chain gene alleles or haplotypes in Swedish patients with multiple sclerosis. J. Neuroimmunol. 31, 141–147 (1991).
Walter, M., Gibson, W., Ebers, G.C. & Cox, D.W. Susceptibility to Multiple Sclerosis associated with the Proximal Heavy Chain Variable Region. J. Clin. Invest. 87, 1266–1273 (1991).
Wood, N. et al. Susceptibility to multiple sclerosis and the immunoglobulin heavy chain variable region. J. Neurol. 242, 677–682 (1995).
Walter, M.S., Gibson, W.T., Ebers, G.C. & Cox, D.W. Susceptibility to multiple sclerosis is associated with the proximal immunoglobulin heavy chain region. J. Clin. Invest. 87, 1266–1273 (1991).
Hashimoto, L., Walter, M., Cox, D. & Ebers, G.C. Immunoglobulin heavy chain variable region polymorphisms in multiple sclerosis susceptibility. J. Neuroimmunol. 44, 77–83 (1993).
Pericak-Vance, M.A. et al. Segregation with markers on chromosome 19q suggest a susceptibility locus for multiple sclerosis (MS). Am. J. Hum. Genef. 55, A199 (1994).
Buetow, K.H. et al. Integrated human genome-wide maps constructed using the CEPH reference panel. Nature Genet. 6, 391–393 (1994).
NIH/CEPH Collaborative Mapping Group.A Comprehensive Genetic Linkage Map of the Human Genome. Science 258, 67–86 (1992).
Risch, N. Linkage strategies for genetically complex traits. III. The effect of marker polymorphism on analysis of affected relative pairs. Am. J. Hum. Genet. 46, 242–253 (1990).
Risch, N. Linkage strategies for genetically complex traits. II. The power of affected relative pairs. Am. J. Hum. Genet. 46, 229–241 (1990).
Risch, N. Exclusion mapping for complex disease. Am. J. Hum. Genet. 53, 185 (1993).
Terwilliger, J.D. & Ott, J. A novel polylocus method for linkage analysis using the lod-score or affected sib-pair method.Genef. Epidemiol. 10, 477–482(1993).
Spielman, R.S., McGinnis, R.E. & Ewens, W.J. Transmission test for linkage disequilibrium: the insulin gene region and insulin-dependent diabetes mellitus (IDDM). Am. J. Hum. Genet. 52, 506–516 (1993).
Author information
Authors and Affiliations
Rights and permissions
About this article
Cite this article
Ebers, G., Kukay, K., Bulman, D. et al. A full genome search in multiple sclerosis. Nat Genet 13, 472–476 (1996). https://doi.org/10.1038/ng0896-472
Received:
Accepted:
Issue Date:
DOI: https://doi.org/10.1038/ng0896-472
This article is cited by
-
Genetic, Immune-Inflammatory, and Oxidative Stress Biomarkers as Predictors for Disability and Disease Progression in Multiple Sclerosis
Molecular Neurobiology (2017)
-
Copy number variation in Y chromosome multicopy genes is linked to a paternal parent-of-origin effect on CNS autoimmune disease in female offspring
Genome Biology (2015)
-
Tumor necrosis factor beta (TNF-β) NcoI polymorphism is associated with multiple sclerosis in Caucasian patients from Southern Brazil independently from HLA-DRB1
Journal of Molecular Neuroscience (2014)
-
Therapies for multiple sclerosis: considerations in the pediatric patient
Nature Reviews Neurology (2011)
-
A Distinct Role of CD4+ Th17- and Th17-Stimulated CD8+ CTL in the Pathogenesis of Type 1 Diabetes and Experimental Autoimmune Encephalomyelitis
Journal of Clinical Immunology (2011)